Code
!wget https://user.it.uu.se/~chrav452/workshop_NN_DL_cnn_data.zip
!unzip workshop_NN_DL_cnn_data.zip
from google.colab import output
output.enable_custom_widget_manager()
!pip install segmentation_models_pytorchTo run this notebook on Google Colab:
!wget https://user.it.uu.se/~chrav452/workshop_NN_DL_cnn_data.zip
!unzip workshop_NN_DL_cnn_data.zip
from google.colab import output
output.enable_custom_widget_manager()
!pip install segmentation_models_pytorchFor this lab, we use the Human White Blood Cell images from Jiangxi Tecom Science Corporation, China.
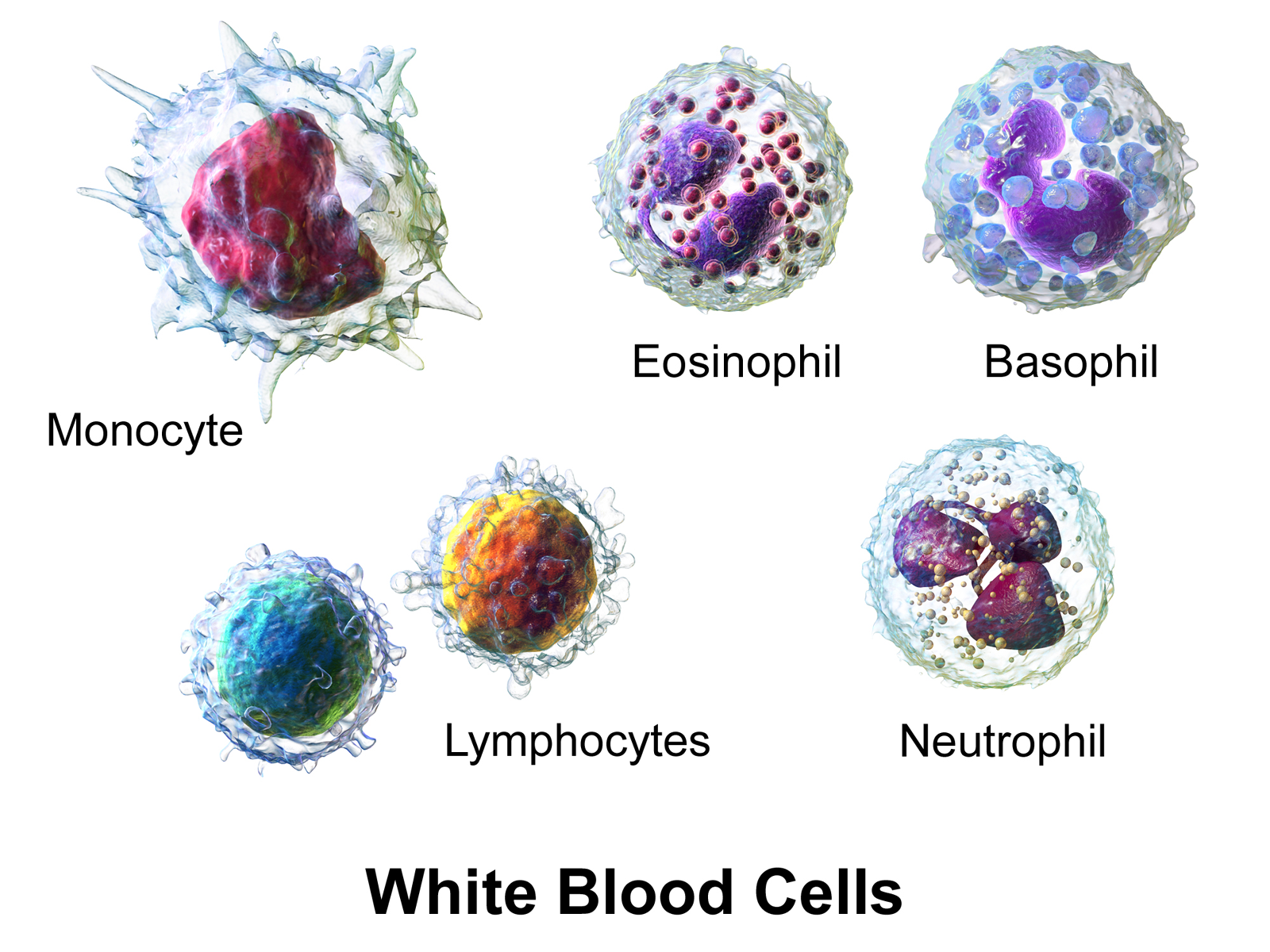
(Illustration from Wikipédia)
The dataset contains three hundred 120×120 RGB images with one blood cell per image, and corresponding segmentation masks. The segmentation mask was manually sketched by domain experts, with the background, cytoplasms and nuclei pixels labelled as 0, 1 and 2 respectively.
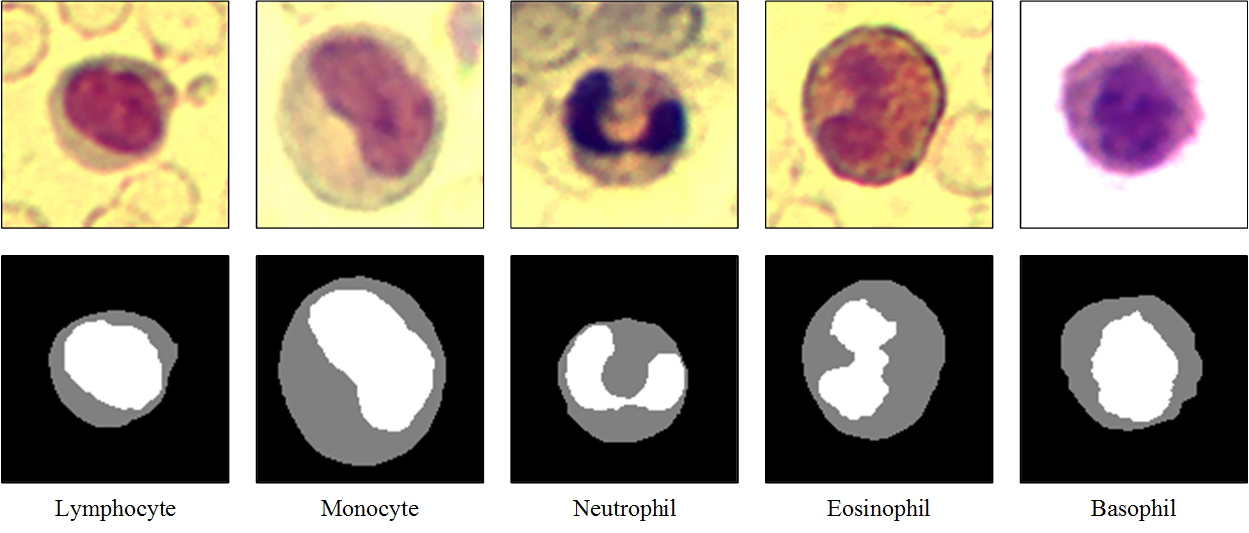
These images and masks are in the data/bloodcells_seg/ folder:
└── data
└── bloodcells_seg
├── masks
│ └── all
└── images
└── allWe want to use convolutional neural networks to do pixel-wise classification of these blood cell images into background / cytoplasm / nuclei.
import random
from pathlib import Path
from typing import Optional, Callable
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import plotly.graph_objects as go
from PIL import Image
from IPython.display import display, clear_output
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader, random_split
from torchvision import transforms
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
print(f"Using device: {device}")We build a small paired dataset that reads RGB images from data/bloodcells_seg/images/all/ and grayscale masks from data/bloodcells_seg/masks/all/. The masks store class indices {0, 1, 2} directly, so we load them as long tensors.
IMG_SIZE = 128
NUM_CLASSES = 3 # background, cytoplasm, nuclei
N_CHANNELS = 3 # RGB
BATCH_SIZE = 8
SEED = 909
class SegDataset(Dataset):
"""Paired image / mask dataset. Assumes filenames match after sorting."""
def __init__(self, image_dir: str, mask_dir: str, img_size: int):
self.image_paths = sorted(Path(image_dir).glob('*'))
self.mask_paths = sorted(Path(mask_dir).glob('*'))
assert len(self.image_paths) == len(self.mask_paths), (
f"Mismatched counts: {len(self.image_paths)} images vs "
f"{len(self.mask_paths)} masks"
)
self.img_size = img_size
self.image_tf = transforms.Compose([
transforms.Resize((img_size, img_size)),
transforms.ToTensor(), # [0,1] RGB, (C,H,W)
])
self.mask_resize = transforms.Resize(
(img_size, img_size),
interpolation=transforms.InterpolationMode.NEAREST,
)
def __len__(self):
return len(self.image_paths)
def __getitem__(self, idx):
img = Image.open(self.image_paths[idx]).convert('RGB')
mask = Image.open(self.mask_paths[idx]).convert('L')
img = self.image_tf(img)
mask = self.mask_resize(mask)
mask = torch.as_tensor(np.array(mask), dtype=torch.long) # (H, W)
return img, mask
dataset = SegDataset(
image_dir='data/bloodcells_seg/images/all',
mask_dir='data/bloodcells_seg/masks/all',
img_size=IMG_SIZE,
)
# Train / development split (80 % train, 20 % dev)
n_total = len(dataset)
n_dev = int(0.2 * n_total)
n_train = n_total - n_dev
train_dataset, dev_dataset = random_split(
dataset, [n_train, n_dev],
generator=torch.Generator().manual_seed(SEED),
)
train_loader = DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True, num_workers=0)
dev_loader = DataLoader(dev_dataset, batch_size=BATCH_SIZE, shuffle=False, num_workers=0)
print(f"Total samples: {n_total}")
print(f"Training samples: {n_train}")
print(f"Development samples: {n_dev}")cMap = ListedColormap(['red', 'lime', 'blue'])
images, masks = next(iter(train_loader))
for i in range(2):
fig, (ax_img, ax_mask) = plt.subplots(1, 2, figsize=(8, 4))
ax_img.imshow(images[i].permute(1, 2, 0).numpy())
ax_img.set_title('Image')
ax_img.axis('off')
ax_mask.imshow(masks[i].numpy(), cmap=cMap, vmin=0, vmax=2)
ax_mask.set_title('Mask')
ax_mask.axis('off')
plt.show()This cell defines a simple live plot used to follow the training and development losses in real time (same LivePlot class as in the classification lab).
from plotly.subplots import make_subplots
class LivePlot():
def __init__(self, prediction_rows=None,
prediction_name: str = "Development predictions"):
self.fig = go.FigureWidget()
self.plot_indices = {}
display(self.fig)
self.limits = [0, 0]
self.current_x = 0
self.grid_figs = {}
if prediction_rows is not None:
self.report_prediction_grid(prediction_name, prediction_rows)
def report(self, name: str, value: float):
try:
plot_index = self.plot_indices[name]
except KeyError:
plot_index = len(self.fig.data)
self.fig.add_scatter(y=[], x=[], name=name)
self.plot_indices[name] = plot_index
self.fig.data[plot_index].y += (value,)
self.fig.data[plot_index].x += (self.current_x,)
def report_prediction_grid(self, name: str, rows):
"""Display rows of (image, truth, prediction) and update in place.
`rows` is a list of dicts: `{'image': (H,W,3) uint8, 'truth': (H,W) int,
'pred': (H,W) int}`. On first call a subplot grid is created; on
subsequent calls the trace data is replaced in place.
"""
n_rows = len(rows)
mask_colorscale = [[0.0, 'red'], [0.5, 'lime'], [1.0, 'blue']]
if name not in self.grid_figs:
fig = make_subplots(
rows=n_rows, cols=3,
column_titles=['Input', 'Ground truth', 'Prediction'],
horizontal_spacing=0.02, vertical_spacing=0.04,
)
for i, row in enumerate(rows):
fig.add_trace(go.Image(z=row['image']), row=i + 1, col=1)
fig.add_trace(go.Heatmap(z=row['truth'], colorscale=mask_colorscale,
zmin=0, zmax=2, showscale=False),
row=i + 1, col=2)
fig.add_trace(go.Heatmap(z=row['pred'], colorscale=mask_colorscale,
zmin=0, zmax=2, showscale=False),
row=i + 1, col=3)
for r in range(1, n_rows + 1):
for c in range(1, 4):
idx = 3 * (r - 1) + c
suffix = '' if idx == 1 else str(idx)
fig.layout[f'xaxis{suffix}'].update(showticklabels=False)
fig.layout[f'yaxis{suffix}'].update(
autorange='reversed', showticklabels=False,
scaleanchor=f'x{suffix}', scaleratio=1,
)
fig.update_layout(height=250 * n_rows, width=800,
margin=dict(l=20, r=20, t=40, b=20),
title=name, showlegend=False)
widget = go.FigureWidget(fig)
display(widget)
self.grid_figs[name] = widget
else:
widget = self.grid_figs[name]
with widget.batch_update():
for i, row in enumerate(rows):
widget.data[3 * i + 0].z = row['image']
widget.data[3 * i + 1].z = row['truth']
widget.data[3 * i + 2].z = row['pred']
def increment(self, n_ticks: int):
self.limits[1] += n_ticks
self.fig.update_layout(xaxis_range=self.limits)
def set_limit(self, n_ticks: int):
self.limits[1] = n_ticks
self.fig.update_layout(xaxis_range=self.limits)
def tick(self, n_ticks: Optional[int] = None):
if n_ticks is None:
n_ticks = 1
self.current_x += n_ticksAfter each epoch we refresh a live grid of (input, ground truth, prediction) rows for a few development examples, so we can follow the training of our network visually. We run it on a fixed batch so the same cells are tracked across epochs.
The helper below turns one batch (images, masks) into a list of rows ready for LivePlot.report_prediction_grid.
def build_prediction_rows(model, batch, device, num_plot: int):
images, masks = batch
model.eval()
with torch.no_grad():
preds = model(images.to(device)).argmax(dim=1).cpu().numpy()
k = min(num_plot, images.size(0))
rows = []
for i in range(k):
img = (images[i].permute(1, 2, 0).numpy() * 255).clip(0, 255).astype(np.uint8)
rows.append({
'image': img,
'truth': masks[i].numpy().astype(np.int32),
'pred': preds[i].astype(np.int32),
})
return rowsBoilerplate training loop. It reports training/development loss after each epoch via LivePlot, and if a plot_batch is supplied it draws the predictions panel for that fixed batch each epoch.
def train(*,
model: nn.Module,
train_loader: DataLoader,
dev_loader: DataLoader,
optimizer: torch.optim.Optimizer,
criterion: nn.Module,
max_epochs: int,
device: Optional[torch.device] = None,
liveplot: Optional[LivePlot] = None,
plot_batch=None,
num_plot: int = 2):
if device is None:
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
for epoch in range(max_epochs):
model.train()
train_loss_acc = 0
train_examples = 0
for x_batch, y_batch in train_loader:
optimizer.zero_grad()
x_batch = x_batch.to(device)
y_hat = model(x_batch)
loss = criterion(y_hat, y_batch.to(device))
loss.backward()
optimizer.step()
train_loss_acc += loss.item() * x_batch.size(0)
train_examples += x_batch.size(0)
model.eval()
dev_loss_acc = 0
dev_examples = 0
with torch.no_grad():
for x_batch, y_batch in dev_loader:
x_batch = x_batch.to(device)
y_hat = model(x_batch)
dev_loss_acc += criterion(y_hat, y_batch.to(device)).item() * x_batch.size(0)
dev_examples += x_batch.size(0)
if liveplot is not None:
liveplot.tick()
liveplot.report("Training loss", train_loss_acc / train_examples)
liveplot.report("Development loss", dev_loss_acc / dev_examples)
if plot_batch is not None and liveplot is not None:
rows = build_prediction_rows(model, plot_batch, device, num_plot)
liveplot.report_prediction_grid("Development predictions", rows)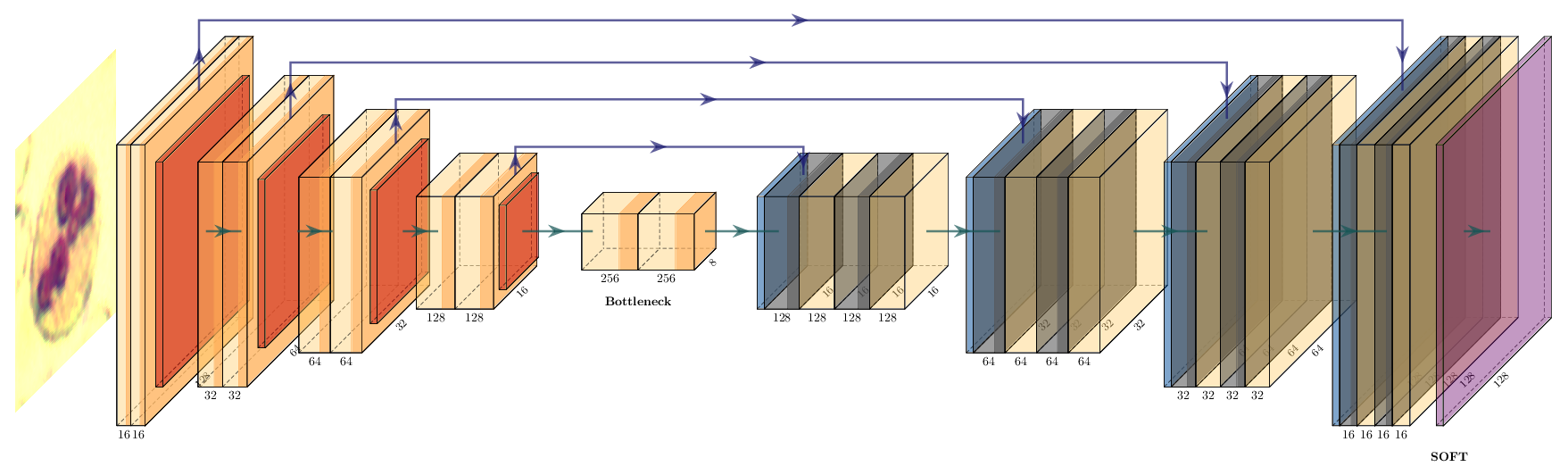
In PyTorch we build the same architecture as a subclass of nn.Module. Each encoder / decoder block is two 3×3 convolutions (with same padding) and an ELU activation, with dropout between the two convolutions.
We use the ELU activation function: the negative values and the smooth transition from negative to positive help us converge faster in a low number of epochs. You can easily try other activations by replacing nn.ELU() with nn.ReLU(), nn.LeakyReLU(), etc.
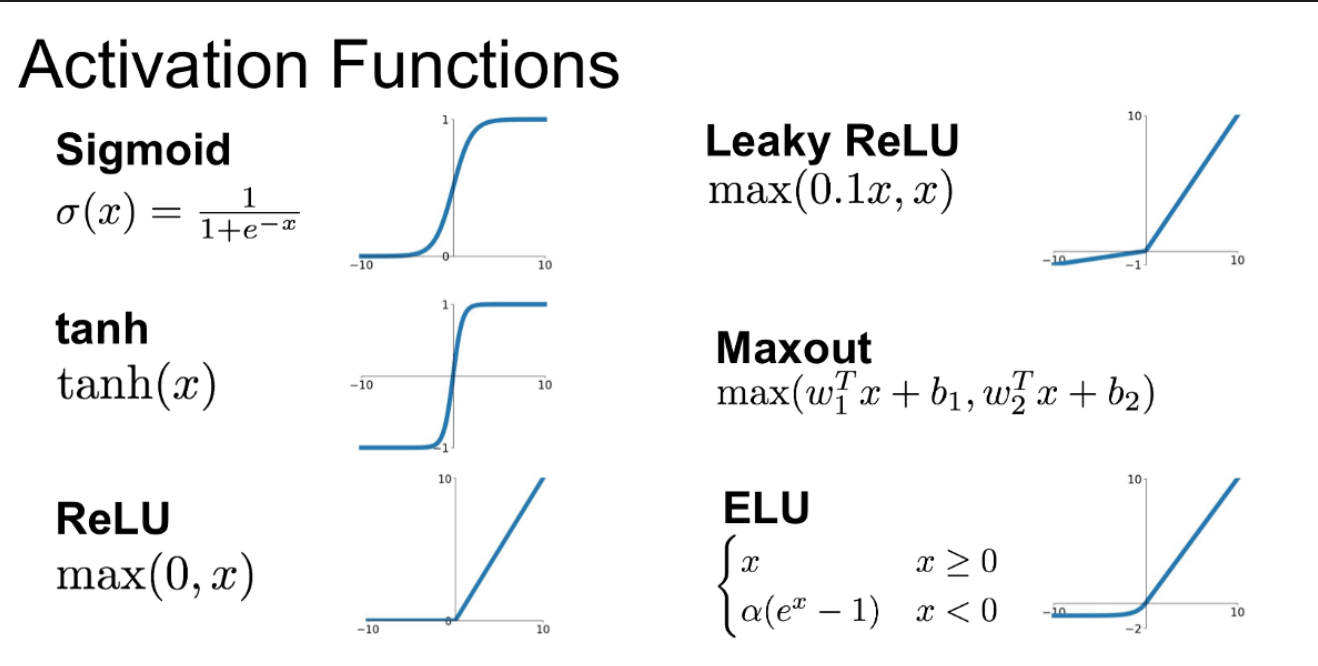

You can visualize differences between activation functions with this online tool from Justin Emery:
from IPython.display import IFrame
IFrame('https://polarisation.github.io/tfjs-activation-functions/', width=860, height=470)class ConvBlock(nn.Module):
"""Two 3×3 'same' convolutions with ELU and dropout in between."""
def __init__(self, in_channels: int, out_channels: int, dropout: float):
super().__init__()
self.block = nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel_size=3, padding=1),
nn.ELU(inplace=True),
nn.Dropout2d(dropout),
nn.Conv2d(out_channels, out_channels, kernel_size=3, padding=1),
nn.ELU(inplace=True),
)
def forward(self, x):
return self.block(x)Note on nn.Dropout2d. Unlike the standard nn.Dropout (which zeroes individual activations), nn.Dropout2d zeroes entire feature-map channels of a 4-D tensor (N, C, H, W). This is the right form of dropout for convolutional layers, where neighbouring spatial activations within a channel are highly correlated.
class UNet(nn.Module):
def __init__(self, in_channels: int = N_CHANNELS, num_classes: int = NUM_CLASSES):
super().__init__()
# Encoder
self.enc1 = ConvBlock(in_channels, 16, 0.1)
self.enc2 = ConvBlock(16, 32, 0.1)
self.enc3 = ConvBlock(32, 64, 0.2)
self.enc4 = ConvBlock(64, 128, 0.2)
self.bottleneck = ConvBlock(128, 256, 0.3)
self.pool = nn.MaxPool2d(2)
# Decoder
self.up4 = nn.ConvTranspose2d(256, 128, kernel_size=2, stride=2)
self.dec4 = ConvBlock(256, 128, 0.2)
self.up3 = nn.ConvTranspose2d(128, 64, kernel_size=2, stride=2)
self.dec3 = ConvBlock(128, 64, 0.2)
self.up2 = nn.ConvTranspose2d(64, 32, kernel_size=2, stride=2)
self.dec2 = ConvBlock(64, 32, 0.1)
self.up1 = nn.ConvTranspose2d(32, 16, kernel_size=2, stride=2)
self.dec1 = ConvBlock(32, 16, 0.1)
# Pixel-wise classifier
self.out_conv = nn.Conv2d(16, num_classes, kernel_size=1)
def forward(self, x):
c1 = self.enc1(x)
c2 = self.enc2(self.pool(c1))
c3 = self.enc3(self.pool(c2))
c4 = self.enc4(self.pool(c3))
c5 = self.bottleneck(self.pool(c4))
d4 = self.dec4(torch.cat([self.up4(c5), c4], dim=1))
d3 = self.dec3(torch.cat([self.up3(d4), c3], dim=1))
d2 = self.dec2(torch.cat([self.up2(d3), c2], dim=1))
d1 = self.dec1(torch.cat([self.up1(d2), c1], dim=1))
return self.out_conv(d1) # logits, shape (N, num_classes, H, W)
model = UNet().to(device)
print(model)
print(f"Total parameters: {sum(p.numel() for p in model.parameters()):,}")The U-Net concatenates upsampled features with the skip connection (torch.cat). Residual networks instead use element-wise addition. Implement a UNetAdd variant that uses addition at each decoder stage instead of concatenation, adjusting the in_channels of each decoder ConvBlock accordingly.
Train both UNet and UNetAdd for 20 epochs and compare their development loss curves and visual predictions. Why does one perform better than the other?
class UNetAdd(nn.Module):
def __init__(self, in_channels: int = N_CHANNELS, num_classes: int = NUM_CLASSES):
super().__init__()
# Encoder — identical to UNet
self.enc1 = ConvBlock(in_channels, 16, 0.1)
self.enc2 = ConvBlock(16, 32, 0.1)
self.enc3 = ConvBlock(32, 64, 0.2)
self.enc4 = ConvBlock(64, 128, 0.2)
self.bottleneck = ConvBlock(128, 256, 0.3)
self.pool = nn.MaxPool2d(2)
# Decoder — YOUR CODE HERE: adjust in_channels since we add, not cat
self.up4 = nn.ConvTranspose2d(256, 128, kernel_size=2, stride=2)
self.dec4 = ConvBlock(???, 128, 0.2)
# ... complete the decoder
self.out_conv = nn.Conv2d(16, num_classes, kernel_size=1)
def forward(self, x):
c1 = self.enc1(x)
c2 = self.enc2(self.pool(c1))
c3 = self.enc3(self.pool(c2))
c4 = self.enc4(self.pool(c3))
c5 = self.bottleneck(self.pool(c4))
# YOUR CODE HERE: use addition instead of torch.cat
d4 = self.dec4(self.up4(c5) + c4)
# ...
return self.out_conv(d1)# source_hidden
# Suggested solution: U-Net with additive skip connections
class UNetAdd(nn.Module):
def __init__(self, in_channels: int = N_CHANNELS, num_classes: int = NUM_CLASSES):
super().__init__()
self.enc1 = ConvBlock(in_channels, 16, 0.1)
self.enc2 = ConvBlock(16, 32, 0.1)
self.enc3 = ConvBlock(32, 64, 0.2)
self.enc4 = ConvBlock(64, 128, 0.2)
self.bottleneck = ConvBlock(128, 256, 0.3)
self.pool = nn.MaxPool2d(2)
# Addition keeps the channel count — no doubling
self.up4 = nn.ConvTranspose2d(256, 128, kernel_size=2, stride=2)
self.dec4 = ConvBlock(128, 128, 0.2)
self.up3 = nn.ConvTranspose2d(128, 64, kernel_size=2, stride=2)
self.dec3 = ConvBlock(64, 64, 0.2)
self.up2 = nn.ConvTranspose2d(64, 32, kernel_size=2, stride=2)
self.dec2 = ConvBlock(32, 32, 0.1)
self.up1 = nn.ConvTranspose2d(32, 16, kernel_size=2, stride=2)
self.dec1 = ConvBlock(16, 16, 0.1)
self.out_conv = nn.Conv2d(16, num_classes, kernel_size=1)
def forward(self, x):
c1 = self.enc1(x)
c2 = self.enc2(self.pool(c1))
c3 = self.enc3(self.pool(c2))
c4 = self.enc4(self.pool(c3))
c5 = self.bottleneck(self.pool(c4))
d4 = self.dec4(self.up4(c5) + c4)
d3 = self.dec3(self.up3(d4) + c3)
d2 = self.dec2(self.up2(d3) + c2)
d1 = self.dec1(self.up1(d2) + c1)
return self.out_conv(d1)
model_add = UNetAdd().to(device)
print(f"UNetAdd parameters: {sum(p.numel() for p in model_add.parameters()):,}")As our ground truth is represented by integer class indices on the mask (0 / 1 / 2) and not one-hot encoded, we use nn.CrossEntropyLoss, which applies log_softmax internally and expects raw logits.
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-4)We pick a fixed development batch so the predictions panel tracks the same cells across epochs.
plot_batch = next(iter(dev_loader))
initial_rows = build_prediction_rows(model, plot_batch, device, num_plot=3)
liveplot = LivePlot(prediction_rows=initial_rows)epochs = 20
liveplot.increment(epochs)
train(model=model, train_loader=train_loader, dev_loader=dev_loader,
optimizer=optimizer, criterion=criterion, max_epochs=epochs,
device=device, liveplot=liveplot,
plot_batch=plot_batch, num_plot=3)rows = build_prediction_rows(model, next(iter(dev_loader)), device, BATCH_SIZE)
liveplot.report_prediction_grid("Final predictions", rows)The standard segmentation metric is IoU per class: \(\text{IoU}_c = |P_c \cap G_c| \,/\, |P_c \cup G_c|\)
Implement per_class_iou returning one score per class. Then compute the pixel-frequency of each class in the training set and display both in a table. Which class has the lowest IoU? Does the class frequency explain it?
def per_class_iou(model, loader, num_classes, device):
# YOUR CODE HERE
pass
class_names = ['background', 'cytoplasm', 'nuclei']
ious = per_class_iou(model, dev_loader, NUM_CLASSES, device)
for name, iou in zip(class_names, ious):
print(f" {name:12s} IoU = {iou:.3f}")
print(f" {'mean':12s} IoU = {np.mean(ious):.3f}")# source_hidden
# Suggested solution: accumulate confusion counts, compute IoU per class
def per_class_iou(model, loader, num_classes, device):
inter = torch.zeros(num_classes)
union = torch.zeros(num_classes)
model.eval()
with torch.no_grad():
for x, y in loader:
preds = model(x.to(device)).argmax(1).cpu()
for cls in range(num_classes):
pred_c = (preds == cls)
true_c = (y == cls)
inter[cls] += (pred_c & true_c).sum()
union[cls] += (pred_c | true_c).sum()
return (inter / union.clamp(min=1)).numpy()
class_names = ['background', 'cytoplasm', 'nuclei']
ious = per_class_iou(model, dev_loader, NUM_CLASSES, device)
for name, iou in zip(class_names, ious):
print(f" {name:12s} IoU = {iou:.3f}")
print(f" {'mean':12s} IoU = {np.mean(ious):.3f}")
# Pixel frequencies in the training set
freq = torch.zeros(NUM_CLASSES)
for _, mask in train_loader:
for cls in range(NUM_CLASSES):
freq[cls] += (mask == cls).sum()
freq = freq / freq.sum()
print()
for name, f in zip(class_names, freq):
print(f" {name:12s} pixel freq = {f:.3f}")You can now try to tune your model by changing the learning rate of the Adam optimizer, the dropout, the batch size, or any other parameter you want.
To simplify the construction of our networks, we can use the segmentation_models_pytorch library. Install it with:
pip install segmentation_models_pytorchThe main features of this library are:
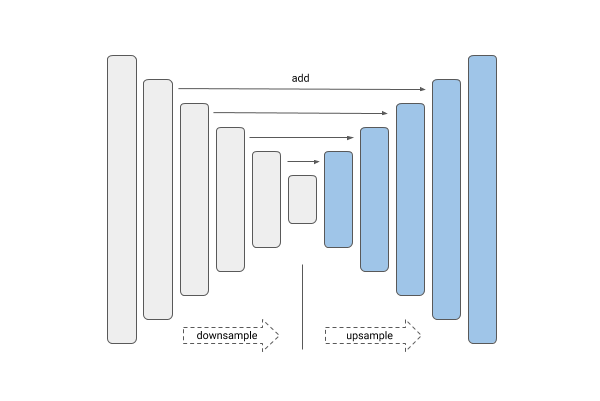
| Unet | Linknet |
|---|---|
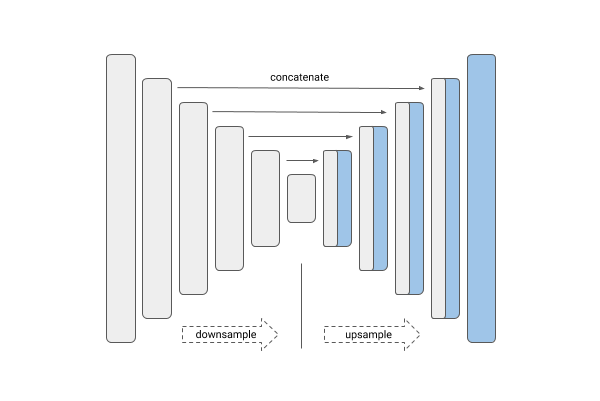 |
 |
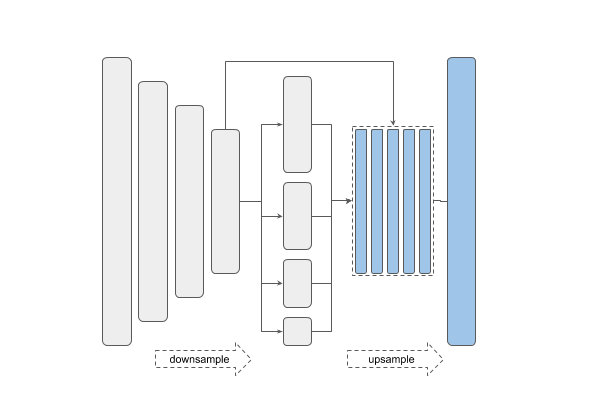
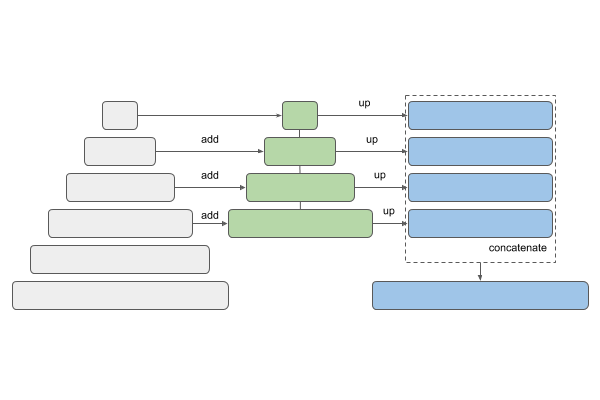
| PSPNet | FPN |
|---|---|
 |
 |
| Type | Names |
|---|---|
| VGG | vgg11 vgg13 vgg16 vgg19 |
| ResNet | resnet18 resnet34 resnet50 resnet101 resnet152 |
| SE-ResNet | se_resnet50 se_resnet101 se_resnet152 |
| ResNeXt | resnext50_32x4d resnext101_32x8d |
| SE-ResNeXt | se_resnext50_32x4d se_resnext101_32x4d |
| SENet154 | senet154 |
| DenseNet | densenet121 densenet169 densenet201 |
| Inception | inceptionv4 inceptionresnetv2 |
| MobileNet | mobilenet_v2 |
| EfficientNet | efficientnet-b0 … efficientnet-b7 |
U-Net and Link-Net are similar and generally used for the kind of segmentation we have here. FPN (Feature Pyramid Networks) are mainly used for object detection, and PSPNet (Pyramid Scene Parsing Networks) are another family of segmentation models — see this overview.
import segmentation_models_pytorch as smp
# Load a U-Net with a VGG16 backbone, trained from scratch
model = smp.Unet(
encoder_name='vgg16',
encoder_weights="imagenet",
in_channels=N_CHANNELS,
classes=NUM_CLASSES,
).to(device)
print(model)
print(f"Total parameters: {sum(p.numel() for p in model.parameters()):,}")criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-4)
initial_rows = build_prediction_rows(model, plot_batch, device, num_plot=2)
liveplot = LivePlot(prediction_rows=initial_rows)epochs = 30
liveplot.increment(epochs)
train(model=model, train_loader=train_loader, dev_loader=dev_loader,
optimizer=optimizer, criterion=criterion, max_epochs=epochs,
device=device, liveplot=liveplot,
plot_batch=plot_batch, num_plot=2)rows = build_prediction_rows(model, next(iter(dev_loader)), device, BATCH_SIZE)
liveplot.report_prediction_grid("Final predictions (smp U-Net / VGG16)", rows)