Code
!wget https://user.it.uu.se/~chrav452/workshop_NN_DL_cnn_data.zip
!unzip workshop_NN_DL_cnn_data.zip
!pip install -q grad-cam
from google.colab import output
output.enable_custom_widget_manager()To run this notebook on Google Colab:
!wget https://user.it.uu.se/~chrav452/workshop_NN_DL_cnn_data.zip
!unzip workshop_NN_DL_cnn_data.zip
!pip install -q grad-cam
from google.colab import output
output.enable_custom_widget_manager()For this lab, we use the image set Human White Blood Cells (BBBC045v1) from the Broad Bioimage Benchmark Collection [Ljosa et al., Nature Methods, 2012].
Using fluorescence staining (Label‐Free Identification of White Blood Cells Using Machine Learning (Nassar et. al)), each blood cell has been classified into one of 5 categories:
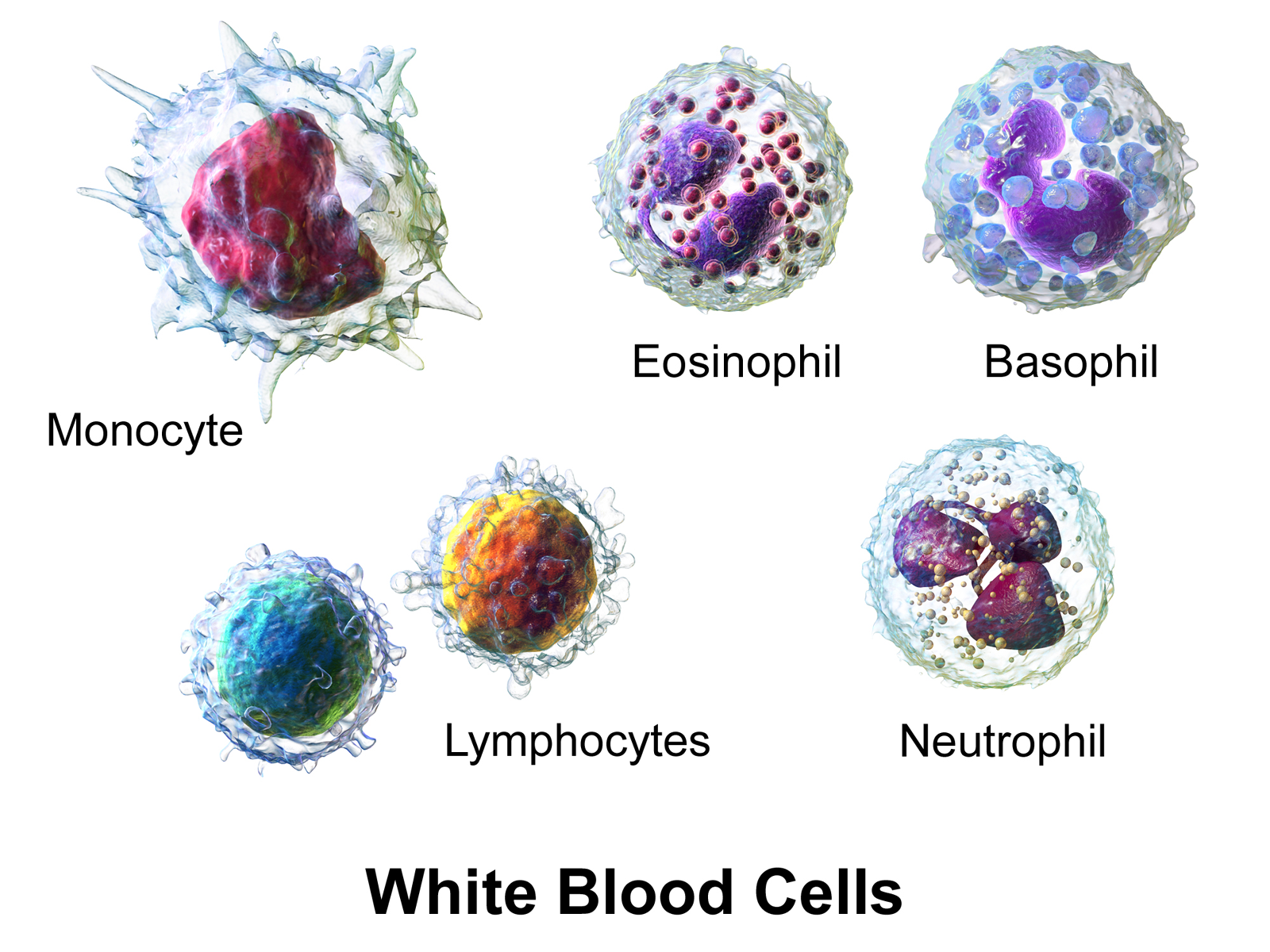
(Illustration from Wikipédia)
For this lab, we only kept Brightfield images and cropped them into small grayscale patches of 32×32 pixels:
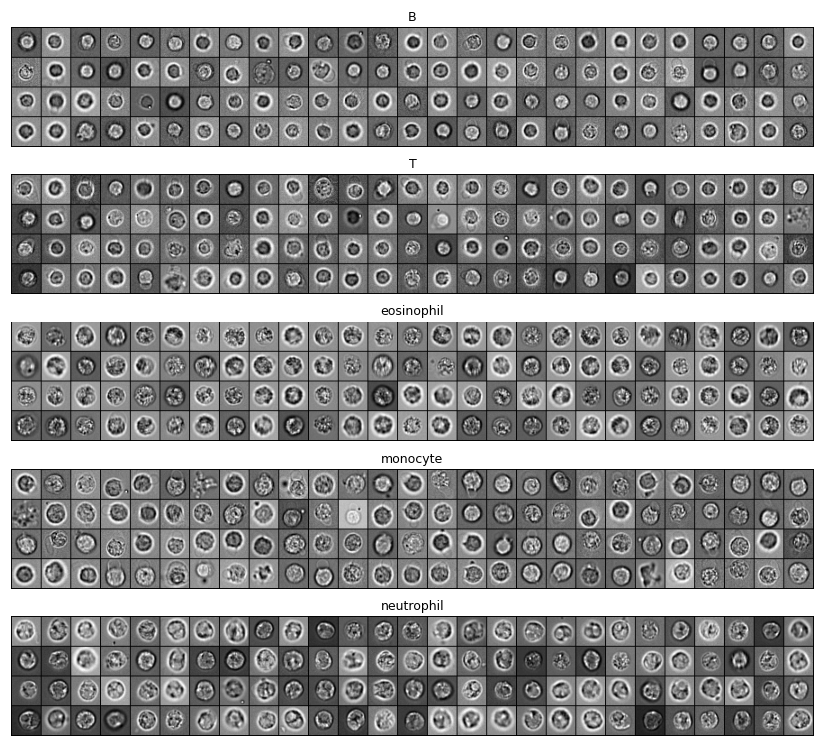

These patches are in the data/bloodcells_small/ folder, split into testing and training sets. In each set, images are split according to their categories:
└── data
└── bloodcells_small
├── test
│ ├── B
│ ├── T
│ ├── eosinophil
│ ├── monocyte
│ └── neutrophil
└── train
├── B
├── T
├── eosinophil
├── monocyte
└── neutrophilOur goal is to use convolutional neural networks to automatically classify blood cells into one of the five categories, using only the 32×32 pixels brightfield images.
import random
import itertools
from typing import Optional
from collections import Counter
import numpy as np
import matplotlib.pyplot as plt
import plotly.graph_objects as go
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
from torchvision import datasets, transforms
from sklearn.metrics import confusion_matrix
# Check for GPU
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
print(f"Using device: {device}")We use ImageFolder and DataLoader to create loaders for the training and testing datasets. ImageFolder reads the class of each image from the directory structure.
We also apply per-sample centering and standardization: each image is shifted to zero mean and scaled to unit standard deviation before being fed to the network.
IMG_SIZE = 32
BATCH_SIZE = 8
NUM_CLASSES = 5
def samplewise_standardize(x: torch.Tensor) -> torch.Tensor:
"""Zero-mean, unit-std normalisation applied independently to each image."""
return (x - x.mean()) / (x.std() + 1e-6)
gray_transform = transforms.Compose([
transforms.Grayscale(),
transforms.Resize((IMG_SIZE, IMG_SIZE)),
transforms.ToTensor(), # [0,1], (C,H,W)
transforms.Lambda(samplewise_standardize),
])
train_dataset = datasets.ImageFolder('data/bloodcells_small/train/', transform=gray_transform)
dev_dataset = datasets.ImageFolder('data/bloodcells_small/test/', transform=gray_transform)
train_loader = DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True, num_workers=0)
dev_loader = DataLoader(dev_dataset, batch_size=BATCH_SIZE, shuffle=False, num_workers=0)
print(f"Classes: {train_dataset.classes}")
print(f"Class to index mapping: {train_dataset.class_to_idx}")
print(f"Training samples: {len(train_dataset)}")
print(f"Testing samples: {len(dev_dataset)}")def class_sizes(dataset):
return dict(Counter(dataset.classes[t] for _, t in dataset.samples))
print("Images per class in training:", class_sizes(train_dataset))
print("Images per class in testing: ", class_sizes(dev_dataset))num_images = 5
for i in range(num_images):
idx = random.randrange(len(train_dataset))
image, label = train_dataset[idx]
img = image.squeeze().numpy() # (H, W) for grayscale display
print(f"Category: {train_dataset.classes[label]} (index={label})")
print(f"Image shape: {img.shape}")
plt.imshow(img, cmap='gray')
plt.axis('off')
plt.show()This cell defines a simple live plot used to follow the training loss, the development loss, and the development confusion matrix in real time.
class LivePlot():
def __init__(self, class_names=None, matrix_name: str = "Confusion matrix (dev)"):
# Loss curves figure (always shown first)
self.fig = go.FigureWidget()
self.plot_indices = {}
display(self.fig)
self.limits = [0, 0]
self.current_x = 0
# Matrix figure placeholder (always shown second)
self.matrix_figs = {}
self._matrix_initialized = {}
self._make_matrix_placeholder(matrix_name, class_names)
def _make_matrix_placeholder(self, name: str, labels=None):
"Create an empty matrix figure that will be filled by `report_matrix`."
if labels is not None:
n = len(labels)
z = np.zeros((n, n))
x_labels = list(labels)
y_labels = list(labels)
else:
z = np.zeros((1, 1))
x_labels = [""]
y_labels = [""]
fig = go.FigureWidget(
data=[go.Heatmap(z=z, x=x_labels, y=y_labels,
colorscale='Blues', zmin=0, zmax=1)]
)
fig.update_layout(
title=name,
xaxis=dict(title='Predicted label', constrain='domain'),
yaxis=dict(title='True label', autorange='reversed',
scaleanchor='x', scaleratio=1),
width=600, height=600,
)
display(fig)
self.matrix_figs[name] = fig
self._matrix_initialized[name] = labels is not None
def report(self, name: str, value: float):
"Report new value for line `name` of the current time step."
try:
plot_index = self.plot_indices[name]
except KeyError:
plot_index = len(self.fig.data)
self.fig.add_scatter(y=[], x=[], name=name)
self.plot_indices[name] = plot_index
self.fig.data[plot_index].y += (value,)
self.fig.data[plot_index].x += (self.current_x,)
def report_matrix(self, name: str, matrix, labels=None):
"Update the heatmap `name` with `matrix`. Creates a placeholder on first call if needed."
matrix = np.asarray(matrix)
n_rows, n_cols = matrix.shape
x_labels = list(labels) if labels is not None else list(range(n_cols))
y_labels = list(labels) if labels is not None else list(range(n_rows))
thresh = float(matrix.max()) / 2. if matrix.max() > 0 else 0.5
annotations = [
dict(x=x_labels[j], y=y_labels[i],
text=f"{100*matrix[i, j]:.0f}%",
showarrow=False,
font=dict(color="white" if matrix[i, j] > thresh else "black"))
for i in range(n_rows) for j in range(n_cols)
]
# If this name was never seen, create a placeholder now.
if name not in self.matrix_figs:
self._make_matrix_placeholder(name, labels)
fig = self.matrix_figs[name]
with fig.batch_update():
# If the placeholder was created without labels, re-set them now
# that we know the real shape.
if not self._matrix_initialized.get(name, False):
fig.data[0].x = x_labels
fig.data[0].y = y_labels
self._matrix_initialized[name] = True
fig.data[0].z = matrix
fig.layout.annotations = annotations
def increment(self, n_ticks: int):
"Increment the currently displayed limits with these many ticks."
self.limits[1] += n_ticks
self.fig.update_layout(xaxis_range=self.limits)
def set_limit(self, n_ticks: int):
"Update the currently displayed to exactly these many ticks."
self.limits[1] = n_ticks
self.fig.update_layout(xaxis_range=self.limits)
def tick(self, n_ticks: Optional[int] = None):
"Update the current time with these many ticks, or 1 tick if omitted."
if n_ticks is None:
n_ticks = 1
self.current_x += n_ticksBoilerplate training loop. It runs max_epochs epochs, reports the training and development loss after each epoch, and (if a LivePlot is supplied) updates a live confusion matrix on the development set.
def train(*,
model: nn.Module,
train_loader: DataLoader,
dev_loader: DataLoader,
optimizer: torch.optim.Optimizer,
criterion: nn.Module,
max_epochs: int,
device: Optional[torch.device] = None,
liveplot: Optional[LivePlot] = None):
if device is None:
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model.to(device)
for epoch in range(max_epochs):
training_loss_acc = 0
training_examples = 0
model.train()
for x_batch, y_batch in train_loader:
optimizer.zero_grad()
x_batch = x_batch.to(device)
y_hat = model(x_batch)
loss = criterion(y_hat, y_batch.to(device))
loss.backward()
optimizer.step()
training_loss_acc += loss.item() * x_batch.size(0)
training_examples += x_batch.size(0)
model.eval()
dev_preds, dev_labels = [], []
with torch.no_grad():
dev_loss_acc = 0
dev_examples = 0
for x_batch, y_batch in dev_loader:
x_batch = x_batch.to(device)
y_hat = model(x_batch)
dev_loss_acc += criterion(y_hat, y_batch.to(device)).item() * x_batch.size(0)
dev_examples += x_batch.size(0)
dev_preds.append(y_hat.argmax(1).cpu().numpy())
dev_labels.append(y_batch.numpy())
if liveplot is not None:
liveplot.tick()
liveplot.report("Training loss", training_loss_acc / training_examples)
liveplot.report("Development loss", dev_loss_acc / dev_examples)
y_pred = np.concatenate(dev_preds)
y_true = np.concatenate(dev_labels)
n_classes = int(max(y_true.max(), y_pred.max())) + 1
cm = confusion_matrix(y_true, y_pred, labels=list(range(n_classes)))
cm_norm = cm.astype('float') / cm.sum(axis=1, keepdims=True).clip(min=1)
class_names = getattr(dev_loader.dataset, 'classes', None)
liveplot.report_matrix("Confusion matrix (dev)", cm_norm, labels=class_names)We can start by building a simple convolutional network with one convolutional layer followed by one max-pooling layer.
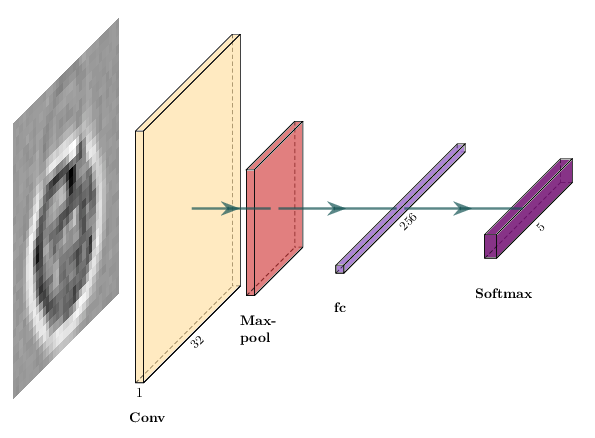
The equivalent in PyTorch is a subclass of nn.Module:
class SimpleCNN(nn.Module):
"""1 conv layer + 1 max-pool + 1 fully connected layer."""
def __init__(self, num_classes=NUM_CLASSES):
super().__init__()
num_filters = 1
filter_size = 2
pool_size = 2
# Conv with 'valid' padding: 32 - 2 + 1 = 31, then pool/2 -> 15
self.conv = nn.Conv2d(1, num_filters, kernel_size=filter_size)
self.pool = nn.MaxPool2d(pool_size)
self.flatten = nn.Flatten()
self.fc = nn.Linear(num_filters * 15 * 15, num_classes)
def forward(self, x):
x = self.conv(x)
x = self.pool(x)
x = self.flatten(x)
return self.fc(x) # logits — CrossEntropyLoss applies softmax internally
model = SimpleCNN().to(device)
print(model)
print(f"Total parameters: {sum(p.numel() for p in model.parameters()):,}")
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters())What is the shape of x after self.conv? After self.pool? Verify your answer by adding print(x.shape) at each step inside forward(), then running a single batch through the model.
Where does the value 15 * 15 in nn.Linear(num_filters * 15 * 15, ...) come from?
images, _ = next(iter(train_loader))
model(images.to(device))# source_hidden
# Suggested solution: shapes through SimpleCNN
print("Input: (B, 1, 32, 32)")
print("After Conv2d(k=2): (B, 1, 31, 31) [32 - 2 + 1 = 31]")
print("After MaxPool2d(2): (B, 1, 15, 15) [31 // 2 = 15]")
print("After Flatten: (B, 225) [1 * 15 * 15 = 225]")We create a fresh LivePlot each time we train a new model so we can keep the history of each run around.
liveplot = LivePlot()epochs = 5
liveplot.increment(epochs)
train(model=model, train_loader=train_loader, dev_loader=dev_loader,
optimizer=optimizer, criterion=criterion, max_epochs=epochs,
device=device, liveplot=liveplot)dev_iter = iter(DataLoader(dev_dataset, batch_size=BATCH_SIZE, shuffle=True))
images, labels = next(dev_iter)
model.eval()
with torch.no_grad():
logits = model(images.to(device))
predictions = logits.argmax(1).cpu().numpy()
print("Predictions: ", predictions)
print("Ground truth: ", labels.numpy())The training loop only reports the loss. Compute the accuracy on the full development set using the trained SimpleCNN. Is the result what you expected given the confusion matrix? A random classifier on 5 balanced classes would achieve 20%.
model.eval()
correct = 0
total = 0
with torch.no_grad():
for x_batch, y_batch in dev_loader:
# YOUR CODE HERE
pass
accuracy = correct / total
print(f"Dev accuracy: {accuracy:.2%}")# source_hidden
# Suggested solution: compute accuracy on the dev set
model.eval()
correct = 0
total = 0
with torch.no_grad():
for x_batch, y_batch in dev_loader:
logits = model(x_batch.to(device))
correct += (logits.argmax(1).cpu() == y_batch).sum().item()
total += y_batch.size(0)
accuracy = correct / total
print(f"Dev accuracy: {accuracy:.2%}")We can now modify our network to improve the accuracy of our classification.
You can add more convolutional and max-pooling layers, and change the number of features in each convolutional layer. For example, two iterations of convolutional layer plus max-pooling, with 16 and 32 features and a kernel size of 3:
nn.Sequential(
nn.Conv2d(1, 16, kernel_size=3),
nn.MaxPool2d(2),
nn.Conv2d(16, 32, kernel_size=3),
nn.MaxPool2d(2),
# ...
)Dropout layers can prevent overfitting. You can add dropout after max-pooling. A dropout of 20% is a good starting point.
nn.Dropout(0.2)Most CNNs use multiple fully-connected layers before the final classifier:
nn.Linear(in_features, 64),
nn.ReLU(),Try adding an activation after convolutional layers (nn.ReLU()), and play with stride and padding (docs):
nn.Conv2d(in_channels, out_channels, kernel_size, stride=2, padding=1)You can change the learning rate of the Adam optimizer:
optimizer = optim.Adam(model.parameters(), lr=1e-4)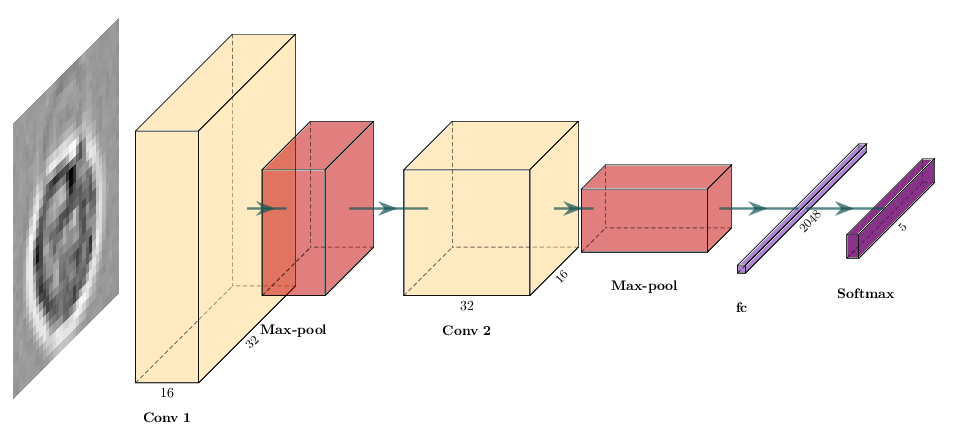
Here we build that model with nn.Sequential, which is a concise alternative to subclassing nn.Module:
num_filters = 16
# After conv(3, valid) + pool(2) twice on a 32x32 input: 32->30->15->13->6
flatten_features = (num_filters * 2) * 6 * 6
model = nn.Sequential(
nn.Conv2d(1, num_filters, kernel_size=3),
nn.ReLU(),
nn.BatchNorm2d(num_filters),
nn.MaxPool2d(2),
nn.Dropout(0.2),
nn.Conv2d(num_filters, num_filters * 2, kernel_size=3),
nn.ReLU(),
nn.MaxPool2d(2),
nn.Dropout(0.2),
nn.Flatten(),
nn.Linear(flatten_features, num_filters * 8),
nn.ReLU(),
nn.Linear(num_filters * 8, NUM_CLASSES),
).to(device)
print(model)
print(f"Total parameters: {sum(p.numel() for p in model.parameters()):,}")
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model.parameters(), lr=1e-4)The advanced model above is provided and ready to train. Before training it, modify it to try the following changes. For each change, state your hypothesis first, then note whether the dev accuracy improves or not.
nn.BatchNorm2d after the second conv block (it is only after the first one right now).nn.Dropout(0.3) layer between the two nn.Linear layers.1e-3 and 5e-5).Keep a copy of each model variant and its dev accuracy so you can compare them after training.
model = nn.Sequential(
# YOUR CODE HERE — modify the advanced CNN above
).to(device)
optimizer = optim.Adam(model.parameters(), lr=1e-4)# source_hidden
# Suggested solution: BatchNorm after the second conv block + dropout
# between the two linear layers
num_filters = 16
flatten_features = (num_filters * 2) * 6 * 6
model = nn.Sequential(
nn.Conv2d(1, num_filters, kernel_size=3),
nn.ReLU(),
nn.BatchNorm2d(num_filters),
nn.MaxPool2d(2),
nn.Dropout(0.2),
nn.Conv2d(num_filters, num_filters * 2, kernel_size=3),
nn.ReLU(),
nn.BatchNorm2d(num_filters * 2),
nn.MaxPool2d(2),
nn.Dropout(0.2),
nn.Flatten(),
nn.Linear(flatten_features, num_filters * 8),
nn.ReLU(),
nn.Dropout(0.3),
nn.Linear(num_filters * 8, NUM_CLASSES),
).to(device)
optimizer = optim.Adam(model.parameters(), lr=1e-4)liveplot = LivePlot()epochs = 5
liveplot.increment(epochs)
train(model=model, train_loader=train_loader, dev_loader=dev_loader,
optimizer=optimizer, criterion=criterion, max_epochs=epochs,
device=device, liveplot=liveplot)VGG16 is a Convolutional Neural Network with five convolutional blocks followed by three fully-connected layers:
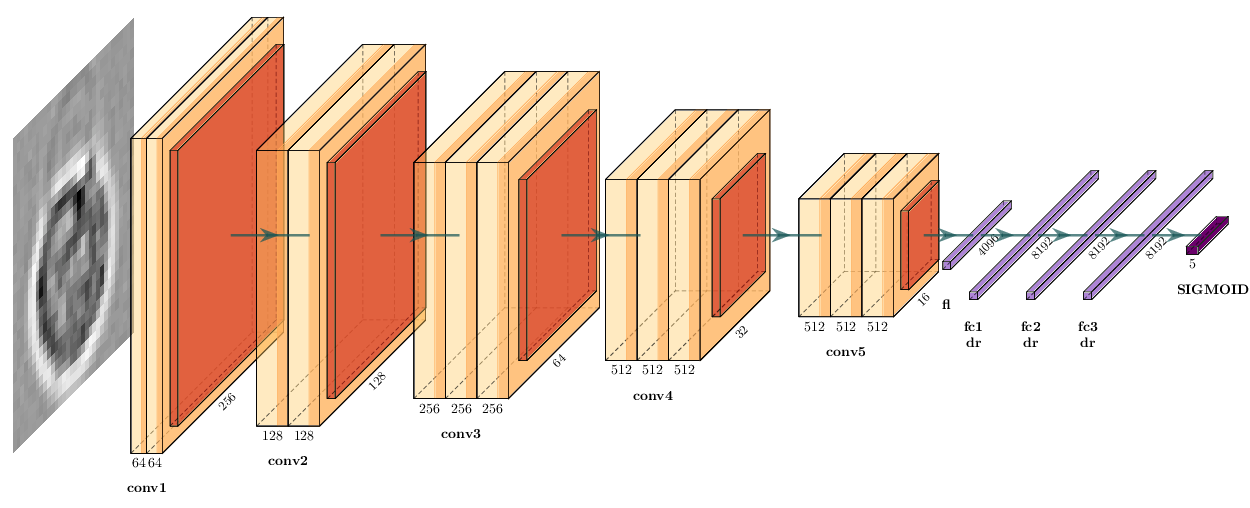
We need to load the data in RGB color mode, as VGG16 expects input with 3 channels:
rgb_transform = transforms.Compose([
transforms.Resize((IMG_SIZE, IMG_SIZE)),
transforms.ToTensor(),
transforms.Lambda(samplewise_standardize),
])
rgb_train = datasets.ImageFolder('data/bloodcells_small/train/', transform=rgb_transform)
rgb_dev = datasets.ImageFolder('data/bloodcells_small/test/', transform=rgb_transform)
rgb_train_loader = DataLoader(rgb_train, batch_size=BATCH_SIZE, shuffle=True, num_workers=2)
rgb_dev_loader = DataLoader(rgb_dev, batch_size=BATCH_SIZE, shuffle=False, num_workers=2)We load VGG16 from torchvision.models (with randomly initialized weights), drop its ImageNet classifier, and replace it with our own. With a 32×32 input, the five max-pool blocks reduce the feature map to 1×1, so we force the adaptive pooling layer to (1, 1) and feed a 512-dim vector into the new classifier head.
from torchvision.models import vgg16, VGG16_Weights
vgg = vgg16(weights=None)
# You can also load pretrained weights with `weights=VGG16_Weights.IMAGENET1K_V1
vgg.avgpool = nn.AdaptiveAvgPool2d((1, 1))
vgg.classifier = nn.Sequential(
nn.Flatten(),
nn.Linear(512, 1024),
nn.ReLU(),
nn.Linear(1024, NUM_CLASSES),
)
vgg = vgg.to(device)
print(vgg)
print(f"Total parameters: {sum(p.numel() for p in vgg.parameters()):,}")
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(vgg.parameters(), lr=1e-4)VGG16 was originally built to work with 224×224 images, while our blood cell crops are only 32×32, which is much smaller. That means we’re pushing the model a bit outside of its intended use here. For now, the goal is just to get a feel for the architecture before we reuse it as an encoder in Lab 2. Because of this mismatch in input size and domain, the pretrained features shouldn’t be expected to transfer perfectly to our data.
liveplot = LivePlot()epochs = 5
liveplot.increment(epochs)
train(model=vgg, train_loader=rgb_train_loader, dev_loader=rgb_dev_loader,
optimizer=optimizer, criterion=criterion, max_epochs=epochs,
device=device, liveplot=liveplot)Earlier we introduced Grad-CAM as a debugging tool: it highlights which regions of the input most influenced the predicted class. We use it here on our trained VGG16 to check whether the network attends to the actual cell or to background / staining artefacts.
We use the pytorch-grad-cam library, which works on any PyTorch model with one extra line:
from pytorch_grad_cam import GradCAM
from pytorch_grad_cam.utils.model_targets import ClassifierOutputTarget
# Our advanced CNN is a single nn.Sequential. Recall the layer indices:
# 0: Conv2d(1, 16) → 30×30
# 3: MaxPool2d → 15×15
# 5: Conv2d(16, 32) → 13×13
# 8: MaxPool2d → 6×6
# We hook the two conv layers and compare their Grad-CAM heatmaps.
cam_conv1 = GradCAM(model=model, target_layers=[model[0]]) # 30×30
cam_conv2 = GradCAM(model=model, target_layers=[model[5]]) # 13×13We pick a few images from the development set and visualise the class-specific heatmap for the predicted class (so we see what justified the model’s decision, right or wrong).
n = 6
model.eval()
fig, axes = plt.subplots(3, n, figsize=(2.2 * n, 6.5))
for col in range(n):
idx = random.randrange(len(dev_dataset))
image, label = dev_dataset[idx] # (1, 32, 32), int
input_tensor = image.unsqueeze(0).to(device)
with torch.no_grad():
pred = model(input_tensor).argmax(1).item()
targets = [ClassifierOutputTarget(pred)]
cam_conv1_map = cam_conv1(input_tensor=input_tensor, targets=targets)[0]
cam_conv2_map = cam_conv2(input_tensor=input_tensor, targets=targets)[0]
# Grayscale image for display
img = image.squeeze(0).cpu().numpy()
img = (img - img.min()) / (img.max() - img.min() + 1e-6)
axes[0, col].imshow(img, cmap='gray')
axes[0, col].set_title(
f"true: {dev_dataset.classes[label]}\npred: {dev_dataset.classes[pred]}",
fontsize=8,
)
axes[0, col].axis('off')
axes[1, col].imshow(img, cmap='gray')
axes[1, col].imshow(cam_conv1_map, cmap='jet', alpha=0.5)
axes[1, col].axis('off')
axes[2, col].imshow(img, cmap='gray')
axes[2, col].imshow(cam_conv2_map, cmap='jet', alpha=0.5)
axes[2, col].axis('off')
axes[1, 0].text(-0.15, 0.5, "Grad-CAM\nmodel[0]\n(30×30 map)",
transform=axes[1, 0].transAxes,
ha='right', va='center', fontsize=8)
axes[2, 0].text(-0.15, 0.5, "Grad-CAM\nmodel[5]\n(13×13 map)",
transform=axes[2, 0].transAxes,
ha='right', va='center', fontsize=8)
plt.tight_layout()
plt.show()::: Grad-CAM is a debugging tool, not an explanation method in any rigorous sense — but it is very effective at catching shortcut learning (model attending to watermarks, scanner artefacts, image borders) before such failures end up in production. :::