Gene set analysis
Single Cell RNA-Seq Analysis
Jennifer Fransson
15-Apr-2026
What is gene set analysis?
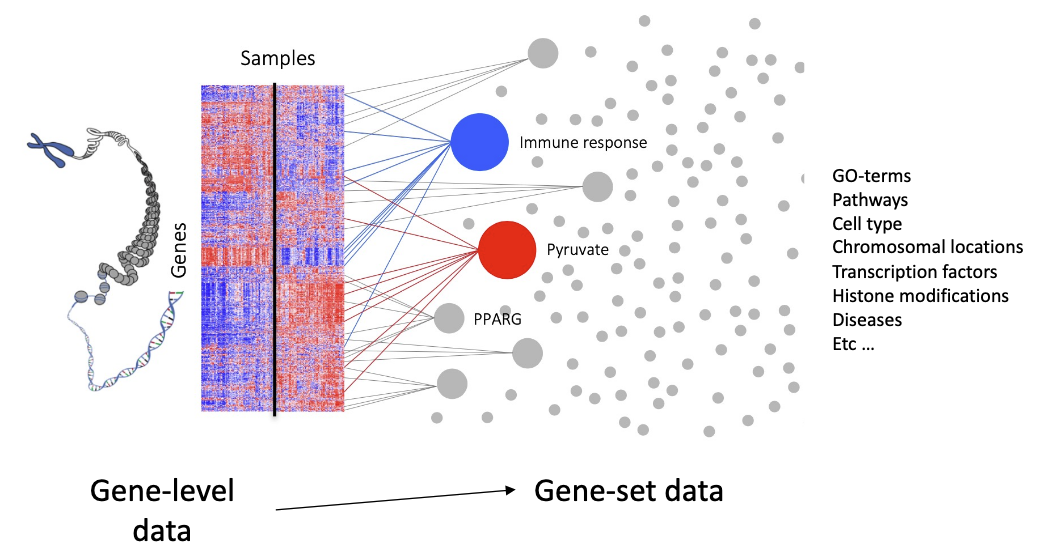
Gene-level data -> Gene set data (Gene set = a list of genes)
We focus on transcriptomics and DGE, but in principle applies to any genome-wide data
Why gene set analysis?
Predict the functional changes of cells based on gene expression
- Make sense of a long list of DEGs
- What is the function of those genes?
- What is the biological consequence of over/under expression of genes?
- Connect your results to pathway activity
- Small differences in many genes may have a bigger impact than large differences in one genes
- Less sensitive to false positive DEGs
Requirements
DE results OR expression data
+
Gene set(s) (list(s) of genes)
+
Statistical test (usually)
Examples of gene sets
- Genes in a pathway
- Genes with shared functions
- Target genes of a transcription factor
- Cell type markers
- Disease-related genes
Where to get gene sets?
- Databases
- Gene Ontology
- KEGG
- Reactome
- MSigDB
- CellMarker
- PanglaoDB
- ChEA
- …
- Literature / previous studies (e.g. list of DE genes, list of transcription factor targets etc)
Different sources vary in level of curation/confidence, interspecies application, number of gene sets, size of gene sets, biases etc
Types of gene set analysis
- Gene set testing
- Find statistically significant enrichment of gene sets in a ranked or non-ranked list of genes
- Activity scoring
- Predict activity of a gene set in individual samples or cells
Gene set testing
DE analysis -> Ranked/unranked list of genes -> Statistical test of enrichment of gene set X
Gene set testing: Overrepresentation analysis (ORA)
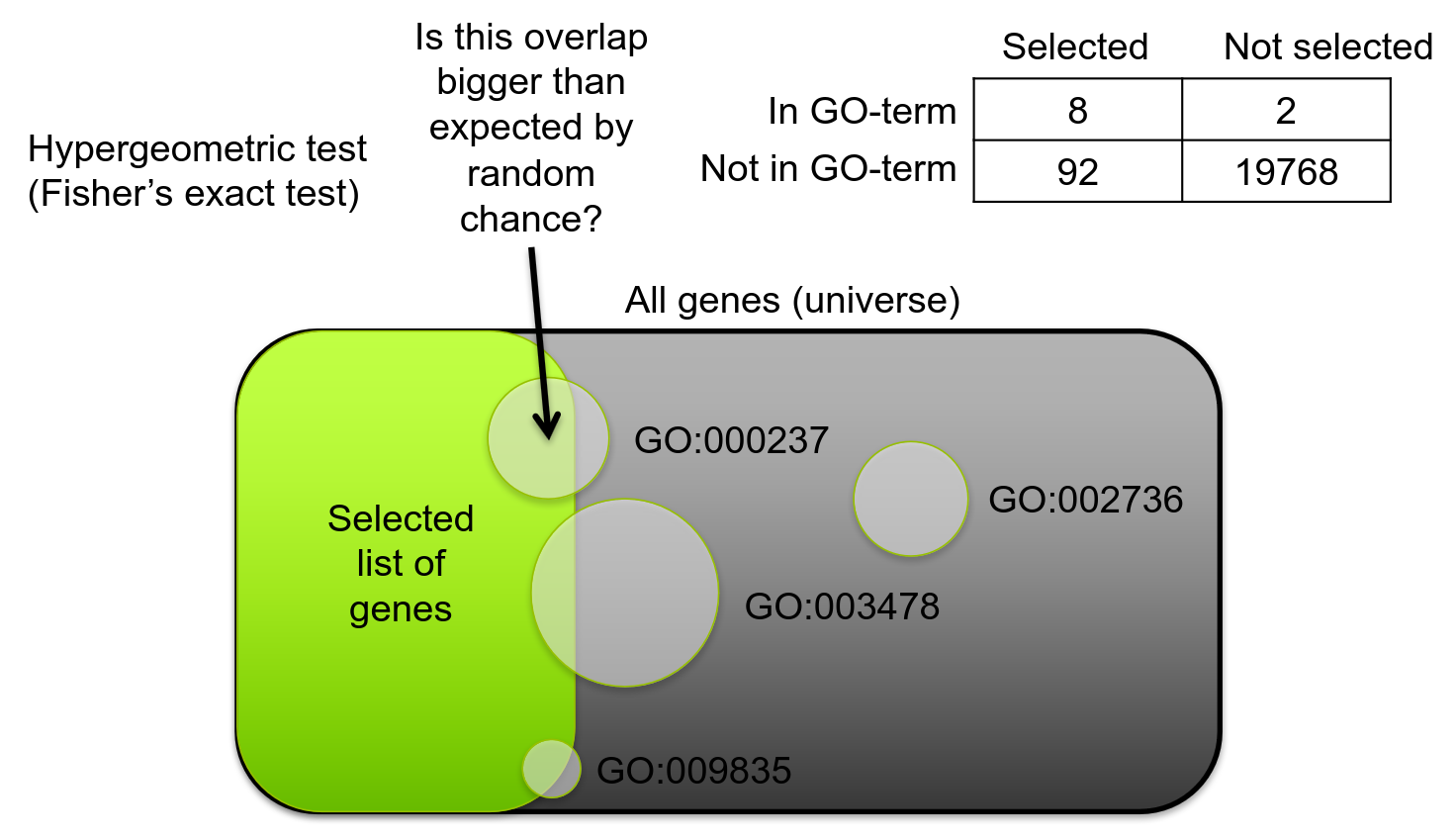
- “Universe” (background) can be all genes or all genes expressed in your cell population
Gene set testing: Overrepresentation analysis (ORA)
- Requires arbitrary cut-off
- Omits actual gene-level statistics
- Selection of genes usually considers both p-value and fold-change
- Computationally fast
- Consider size of overlap in small gene sets!
Gene set testing: Gene set enrichment analysis (GSEA)
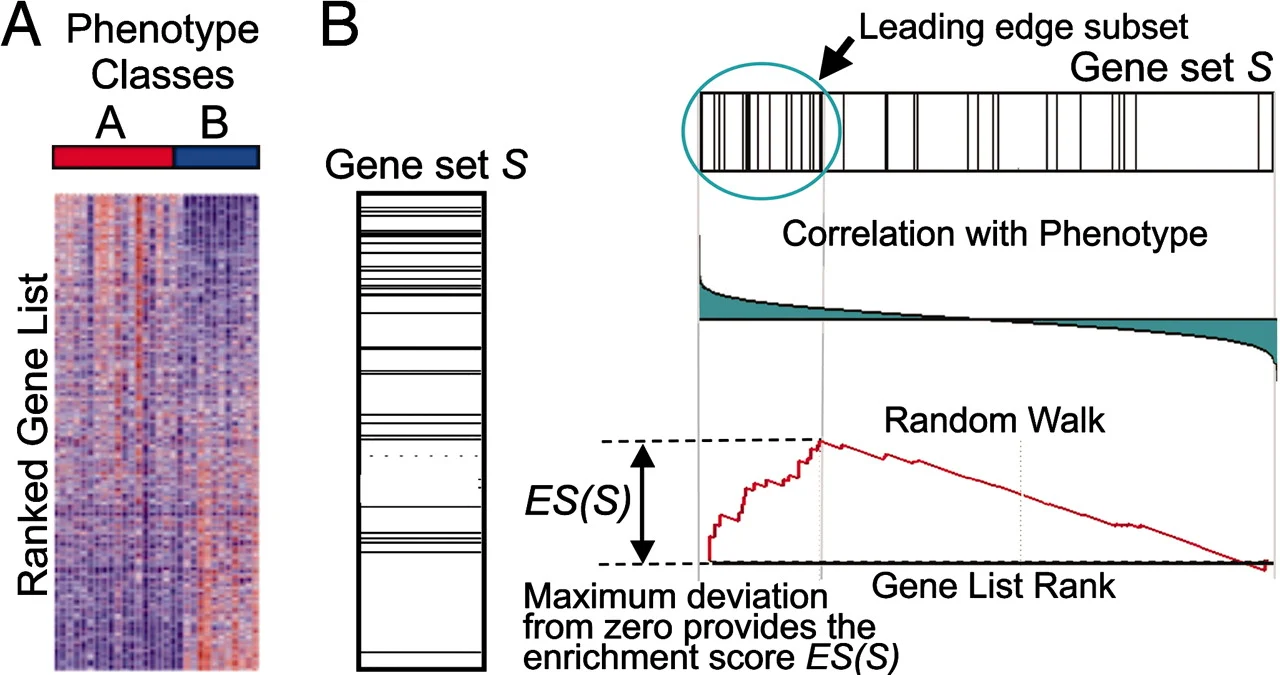
Subramanian et al. (2005)
Gene set testing: Gene set enrichment analysis (GSEA)
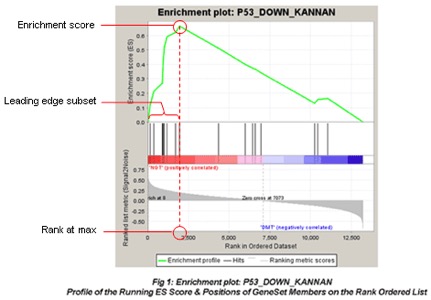
- Enrichment score (ES)
- Normalized enrichment score (NES, corrected for gene set size)
- No need for cut-offs
- Takes gene-level stats into account
- Genes must be ranked according to one variable (usually either log2(FC) or sign(FC) * -log10(p))
- More sensitive to subtle changes
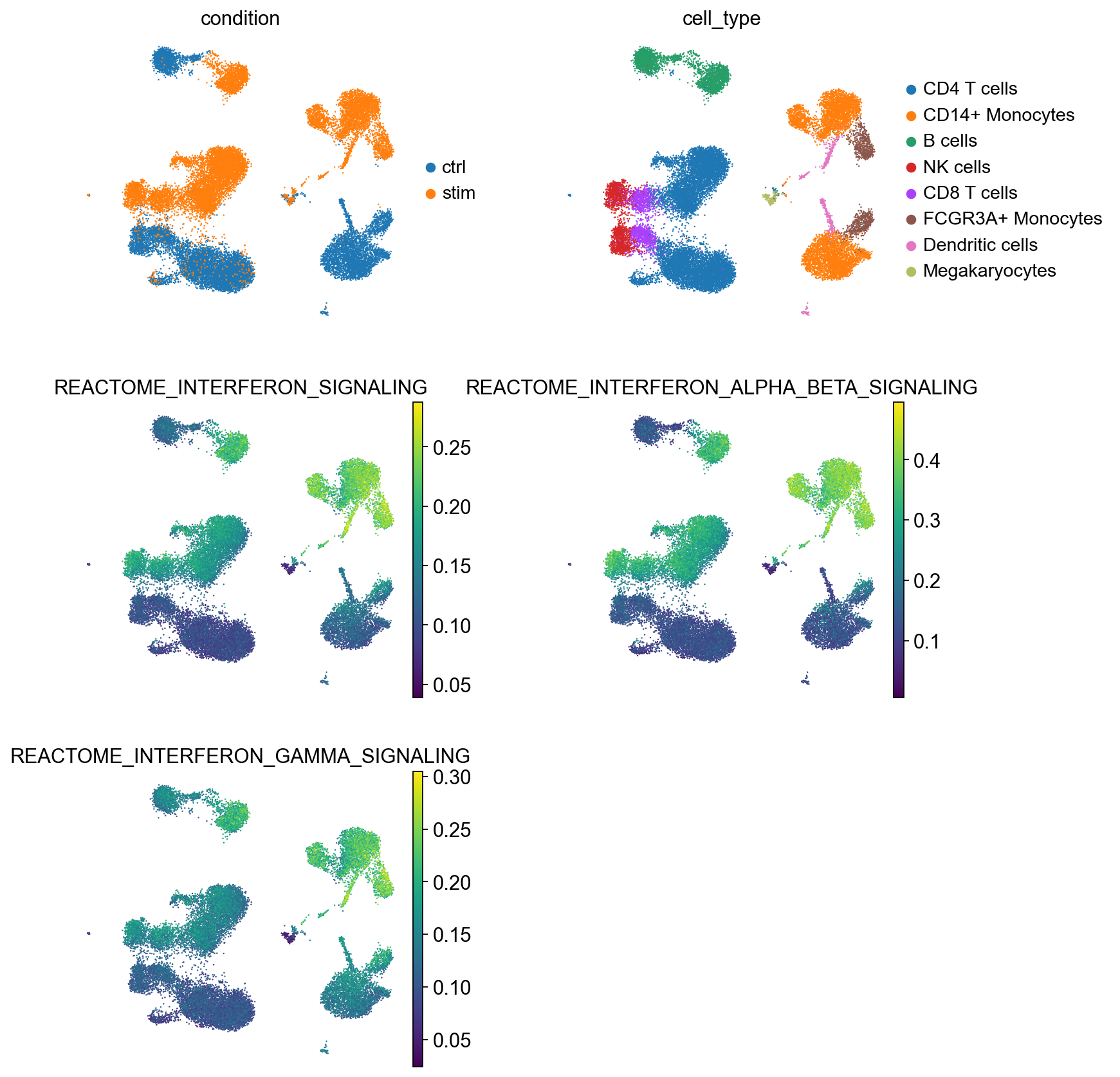
Activity scoring
- For each gene set, calculate score for each sample/cell
- Based on gene expression (not DE)
- Statistical test can then be performed between groups or against a continuous variable
Activity scoring methods (examples)
- Combined z-score
- Control-corrected average expression
- AddModuleScore (Seurat), score_genes (scanpy)
- AUCell (Aibar et al., 2017)
- UCell (Andreatta & Carmona, 2021)
- Vision (DeTomaso et al., 2019)
- Pagoda2 (Lake et al., 2018)
- SCSE (Pont et al., 2019)
- JASMINE (Noureen et al., 2022)
- PROGENy (pathway activity) (Schubert et al., 2018)
- ssGSEA (Barbie et al., 2009)
- GSVA (Hänzelmann et al., 2013)
Benchmarking
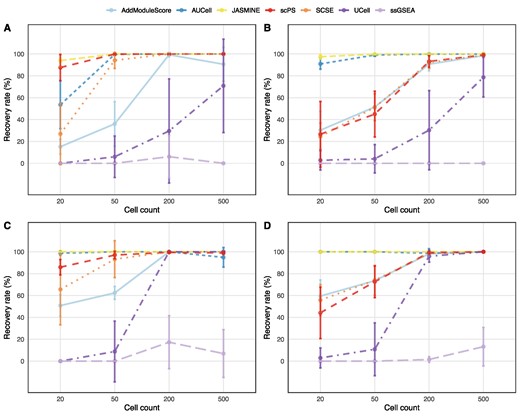
- ssGSEA and GSVA usually under-perform
(Zhang et al., 2020) (Noureen et al., 2022) (Wang & Thakar, 2024)
Considerations
- Enrichment ≠ function
- Activating vs inhibiting genes
- Bias in curation - highly researched topics will be over-represented
- Gene set names can be misleading
- Specific vs general gene sets
- Multifunctional genes
- Translation between different gene IDs
- Protein-based databases
- Databases change
- Curation is organism-specific
- Critical evaluation is required!
References
Aibar, S., González-Blas, C. B., Moerman, T., Huynh-Thu, V. A., Imrichova, H., Hulselmans, G., Rambow, F., Marine, J.-C., Geurts, P., Aerts, J., Oord, J. van den, Atak, Z. K., Wouters, J., & Aerts, S. (2017). SCENIC: Single-cell regulatory network inference and clustering. Nature Methods, 14(11), 1083–1086. https://doi.org/10.1038/nmeth.4463
Andreatta, M., & Carmona, S. J. (2021). UCell: Robust and scalable single-cell gene signature scoring. Computational and Structural Biotechnology Journal, 19, 3796–3798. https://doi.org/https://doi.org/10.1016/j.csbj.2021.06.043
Barbie, D. A., Tamayo, P., Boehm, J. S., Kim, S. Y., Moody, S. E., Dunn, I. F., Schinzel, A. C., Sandy, P., Meylan, E., Scholl, C., Fröhling, S., Chan, E. M., Sos, M. L., Michel, K., Mermel, C., Silver, S. J., Weir, B. A., Reiling, J. H., Sheng, Q., … Hahn, W. C. (2009). Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature, 462(7269), 108–112.
DeTomaso, D., Jones, M. G., Subramaniam, M., Ashuach, T., Ye, C. J., & Yosef, N. (2019). Functional interpretation of single cell similarity maps. Nature Communications, 10(1), 4376. https://doi.org/10.1038/s41467-019-12235-0
Hänzelmann, S., Castelo, R., & Guinney, J. (2013). GSVA: Gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics, 14(1), 7.
Lake, B. B., Chen, S., Sos, B. C., Fan, J., Kaeser, G. E., Yung, Y. C., Duong, T. E., Gao, D., Chun, J., Kharchenko, P. V., & Zhang, K. (2018). Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol., 36(1), 70–80.
Noureen, N., Ye, Z., Chen, Y., Wang, X., & Zheng, S. (2022). Signature-scoring methods developed for bulk samples are not adequate for cancer single-cell RNA sequencing data. Elife, 11(e71994).
Pont, F., Tosolini, M., & Fournié, J. J. (2019). Single-Cell signature explorer for comprehensive visualization of single cell signatures across scRNA-seq datasets. Nucleic Acids Res., 47(21), e133.
Schubert, M., Klinger, B., Klünemann, M., Sieber, A., Uhlitz, F., Sauer, S., Garnett, M. J., Blüthgen, N., & Saez-Rodriguez, J. (2018). Perturbation-response genes reveal signaling footprints in cancer gene expression. Nature Communications, 9(1), 20. https://doi.org/10.1038/s41467-017-02391-6
Subramanian, A., Tamayo, P., Mootha, V. K., Mukherjee, S., Ebert, B. L., Gillette, M. A., Paulovich, A., Pomeroy, S. L., Golub, T. R., Lander, E. S., et al. (2005). Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences, 102(43), 15545–15550. https://www.pnas.org/doi/abs/10.1073/pnas.0506580102
Wang, R. H., & Thakar, J. (2024). Comparative analysis of single-cell pathway scoring methods and a novel approach. NAR Genom. Bioinform., 6(3), lqae124.
Zhang, Y., Ma, Y., Huang, Y., Zhang, Y., Jiang, Q., Zhou, M., & Su, J. (2020). Benchmarking algorithms for pathway activity transformation of single-cell RNA-seq data. Comput. Struct. Biotechnol. J., 18, 2953–2961.
Acknowledgements
Adapted from previous presentations by Leif Wigge, Paulo Czarnewski and Roy Francis.